Your Ideal Biohealth Career Awaits
Wisconsin’s Biohealth Industry is Booming
Whether you’re just launching your career or you’ve been in the industry for years, you’ll find incredible opportunities in our vast network of organizations.







Explore Biohealth Opportunities by Industry
Wisconsin’s Latest Biohealth News
Microbial Discovery Group expands into third facility
WiBxTEMPO Madison – Board Readiness Workshop
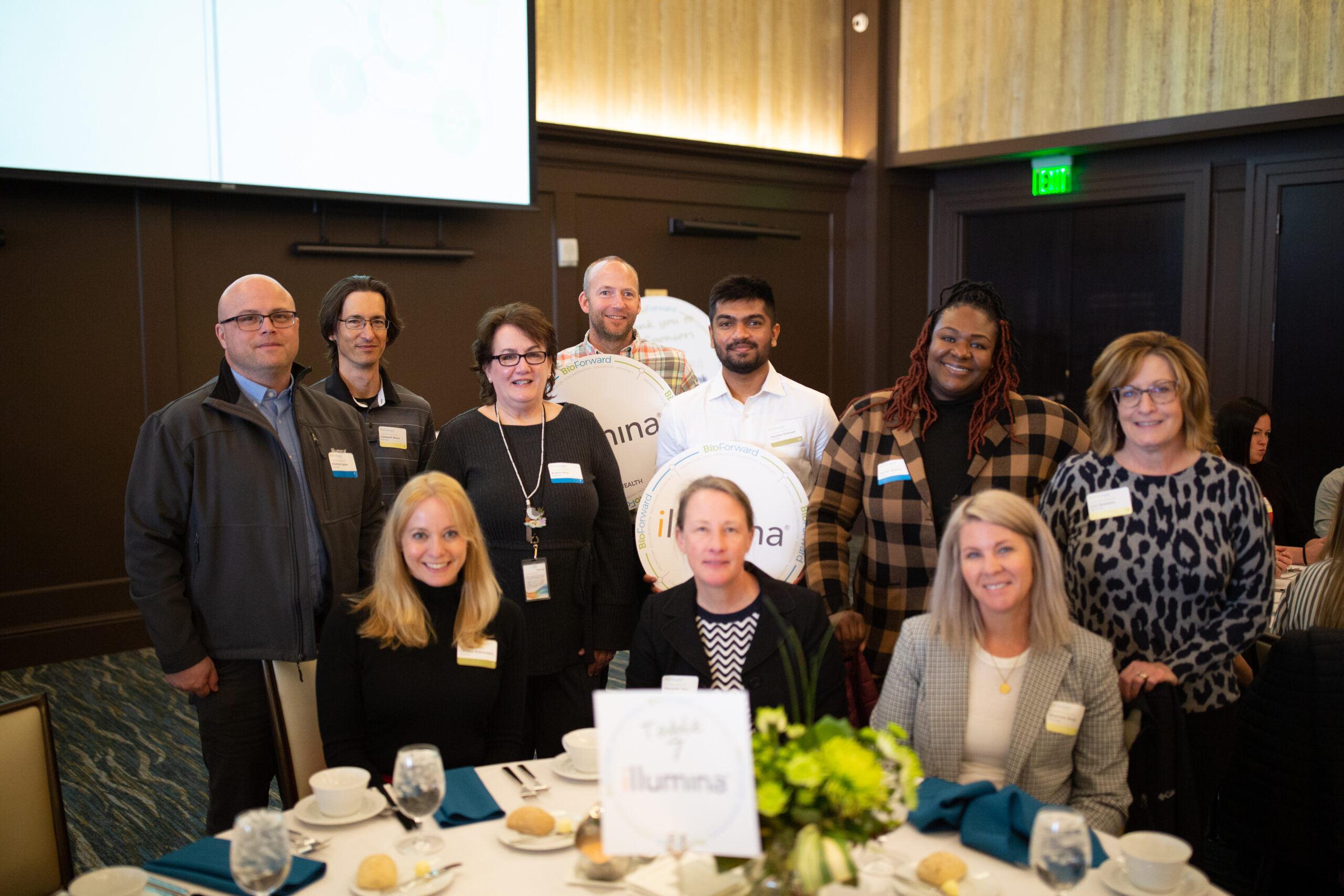
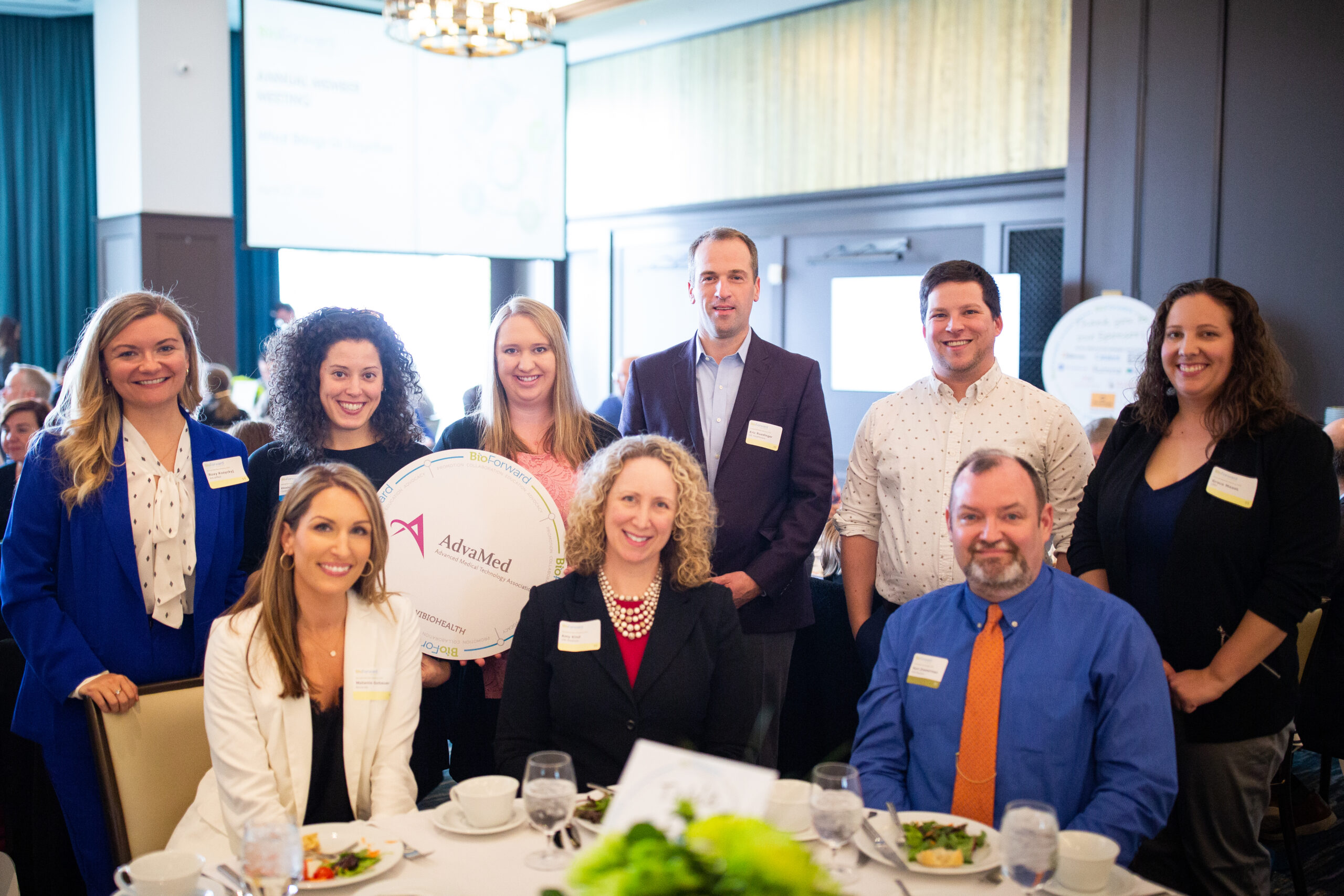
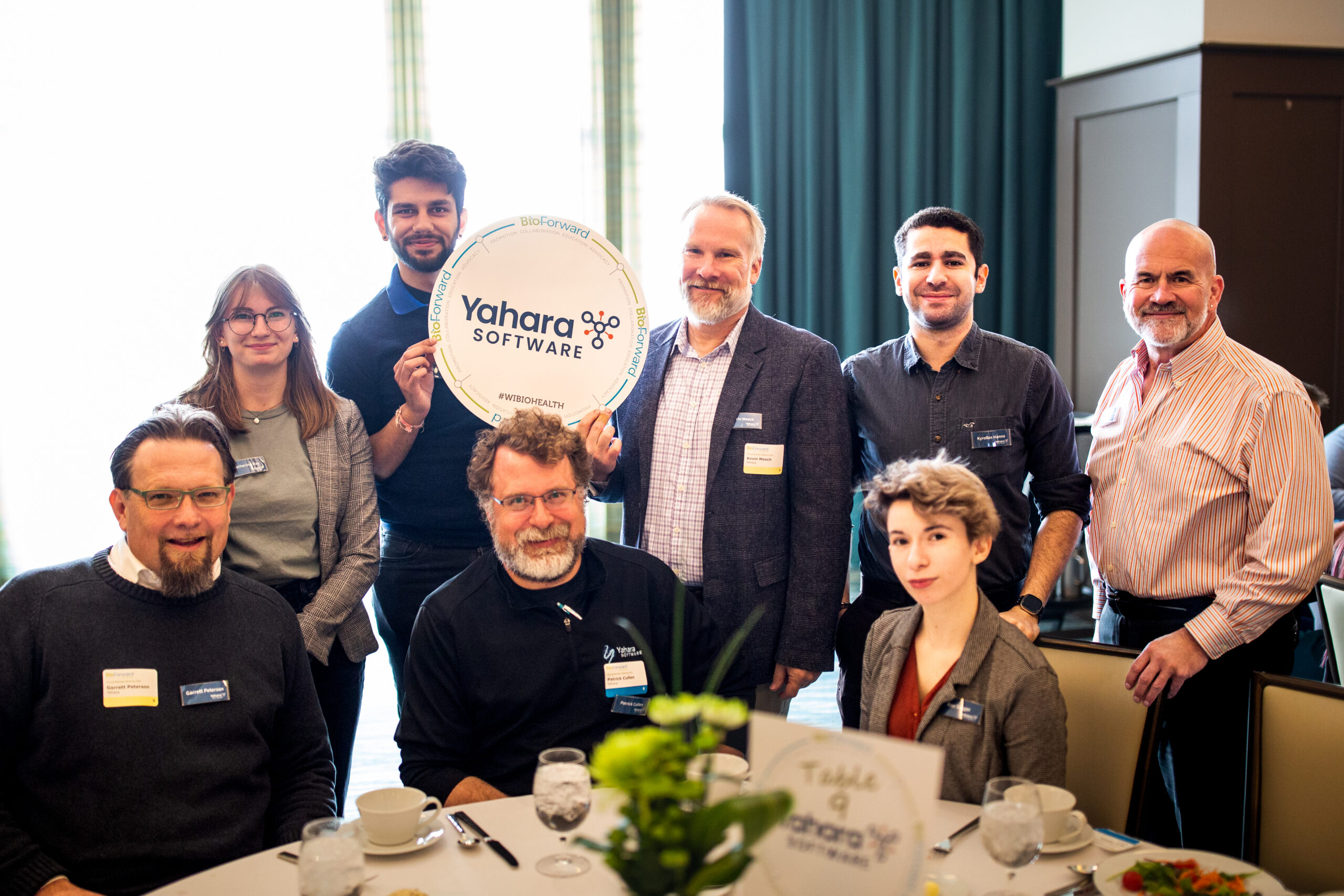
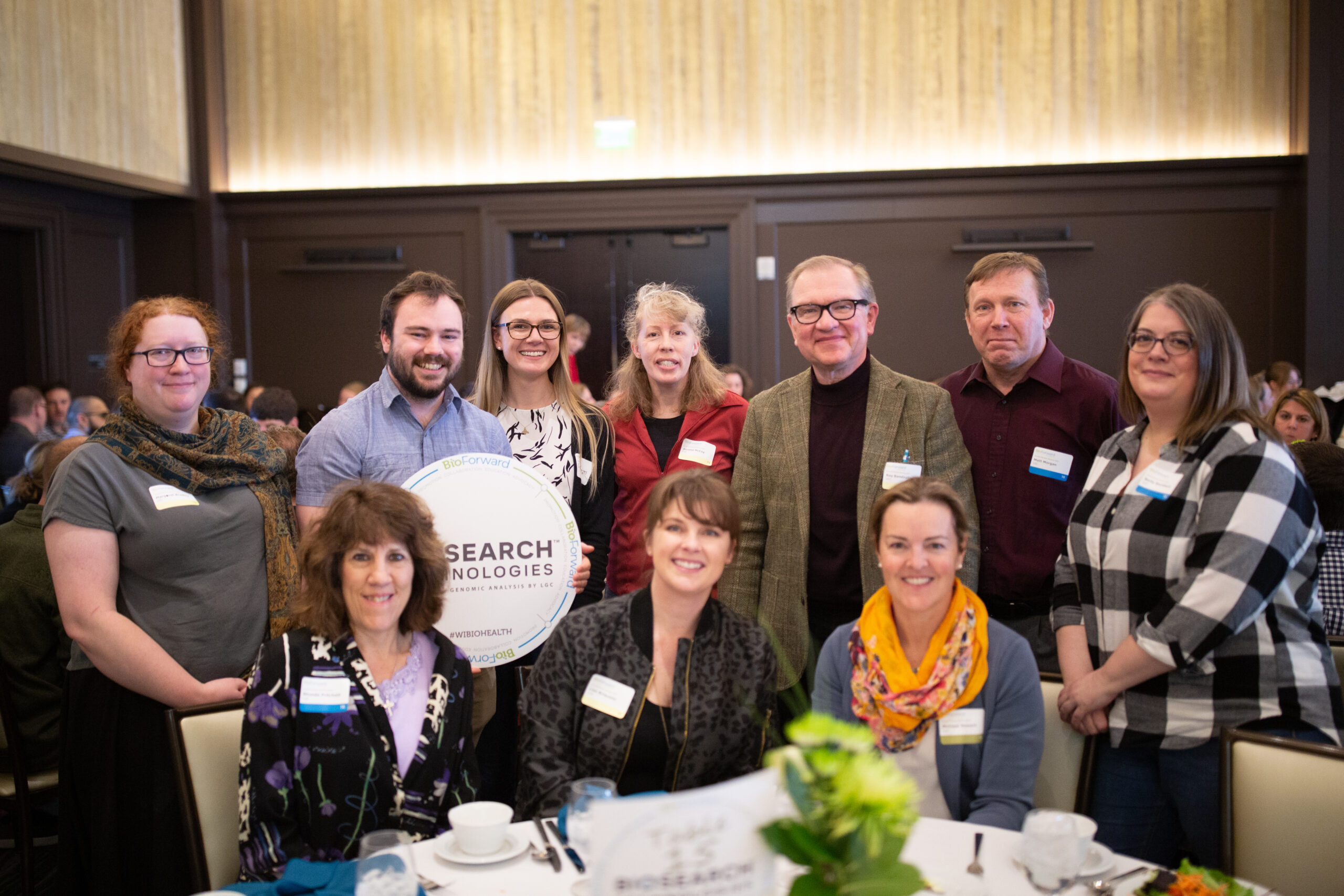
Meet a Few of Wisconsin’s Biohealth Leaders
Our member companies are shaping the Wisconsin biohealth industry by exploring new technologies that improve and save people’s lives, while also making Wisconsin a great place to live, work and play.
About Talent Initiative & BioForward Wisconsin
The Talent Initiative is an effort of BioForward Wisconsin, which offers services and resources to promote the growth and influence of Wisconsin’s bioindustry throughout the U.S. and the world. BioForward is the only Wisconsin organization to represent over two hundred member companies across a variety of biohealth industries, research institutions, and service providers. BioForward offers its members collaborative assistance, educational and networking events, legislative advocacy, and resource investment in key industry initiatives. Learn more about BioForward and its members at bioforward.org.